
Enabling High Throughput 3D Cell Culture Using Automation
Introduction
In many applications, 3D cell culture models have been established as a more accurate research model than 2D cultures. This is particularly important in research where in vivo biological relevance is critical. The most common 3D culture models are spheroids (3D cell aggregates). They can be generated from a wide range of cell types and methods. Spheroids form spontaneously and some examples of multi-cellular spheroids include embryoid bodies, mammospheres, tumor spheroids, hepatospheres and neurospheres.
Applications for 3D Culture – Cancer Research
One area where 3D culture models can be very useful is in cancer research where 3D culture models can be used to predict and model drug responses for drug compound screening and toxicity testing. 3D culture is particularly important here as there is significant evidence that 3D models frequently respond differently to compounds than 2D models. These 3D culture models can create systems to mimic healthy cells or diseased cells and by using these systems, researchers can evaluate cellular responses to different drug candidates early in development. Researchers can also culture tumor cells and conduct tumor invasion assays with tumor spheroids to visualize the invasion of cancer cells and also screen inhibitors. For more information about 3D Cell Culture and research applications, please see, “Is Now the Time to Move Your Research to 3D Cell Culture? Key Considerations and New Tools to Ensure Success.”
Challenges to Implementation
While the benefits of implementing 3D culture are clear, conducting 3D culture manually is inconsistent, time consuming and labor intensive. It is only recently that implementation has grown due to new tools and technologies designed to make 3D culture more efficient and high throughput. High throughput is key in screening large numbers of drug compounds in a reasonable amount of time. Thus you need a way to quickly and efficiently plate large numbers of cell suspensions, and of course it isn’t sufficient just to create the cultures, you must have an efficient way to conduct the analysis as well.
The Solution – Automation
Why automate
Automation solves many challenges associated with implementing 3D culture systems and high throughput screening. Specific benefits include:
Improves consistency, reduces errors and eliminates user-to-user variability
- Automated plating provides spheroids with consistent size and shape for analysis.
- Data can be tracked through each automated step of your process, thus reducing the chance of error. In addition, sample ID and other analytical data will be tracked and recorded from sample input to the end result.
High throughput, thus reducing labor requirements and time at the bench
- Automation allows scientists to significantly reduce labor intensive and time-consuming work. With systems like the Biomek workstation, cell seeding, feeding, compound treatment and sample prep can all be achieved through automation.
- Flow cytometry provides more comprehensive cell-level data that would otherwise be difficult to obtain using manual methods due to the complex sample prep.
- With drug compound screening there can be thousands of compounds that one might want to screen against a certain type of tumor. The vast numbers involved in this kind of task makes manual culturing unrealistic. Automation allows for more comprehensive screening, which provides more possible targets faster and with higher consistency.
Automation Provides the Necessary High Throughput
A recent webinar presented by Beckman Coulter and 3D Biomatrix, “Automated 3D Cell Culture and Screening by Imaging and Flow Cytometry,” demonstrated the ability to automate 3D plating, culture and compound screening with cancer spheroids and the follow up analysis. In the study their goal was to automate the entire process from plating cells to spheroid treatment and apoptosis analysis using both imaging and flow cytometry. I have included here a brief summary of the successful study.
The study
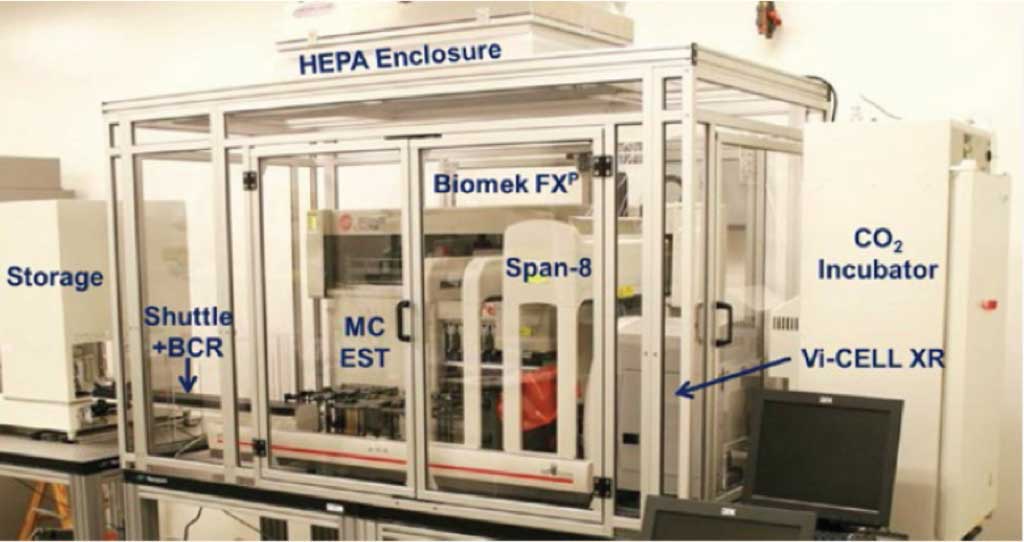
To fully automate a 3D culture process, they employed Beckman Coulter’s Biomek FXP Workstation. Although there are different configurations available, for this project a 96-channel head and a Span-8 pipetting mechanism was used with a HEPA-filter enclosure to ensure a sterile environment. Cells were plated using 3D Biomatrix’s Perfecta3D 96-well hanging drop plates. These plates utilize the hanging drop method in culturing spheroids and provide one of the plate options compatible with automation. 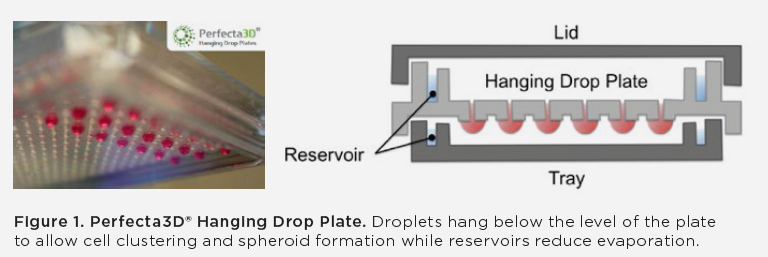
To use the hanging drop method, the cell suspension must be dispensed into the well where the surface tension of the media causes a hanging drop to form. Then cells sink to the bottom and aggregate, which results in spheroid formation within 1-5 days.

Day 0
On day 0 using the Biomek FXP, 4,000 HCT116 colon cancer cells were plated in 40 μl of media using the Perfecta3D 96-well hanging drop plates. In addition 0.25% polyvinyl alcohol was added to accelerate spheroid formation. The cells were plated column-wise using 96-channel enhanced selective tip (EST) pipetting. The goal for the automation was to achieve consistent cell distribution, no lost drops and no contamination, as antibiotics were not used in the culture.
The results showed that a full plate could be processed in less than 8 minutes. Key issues that were addressed to make automation compatible with hanging drop spheroid culture included, ensuring that the pipette positioning was compatible. Reducing dead volume through the use of enhanced multichannel selective tip pipetting, which allowed access to a smaller reservoir and therefore eliminated the need for excess volume. Lastly, slowing the speed to ensure that hanging drops weren’t lost due to being jarred. As a result of these adjustments, no drops were lost and dead volume was reduced to < 1 mL from greater than >20 mL.
Day 3
Robust spheroids were generated by day 3. Prior to compound addition, the spheroids were checked for consistency across wells, which is critical in compound screening. After being analyzed, it was determined that the spheroids showed consistent size and circularity. Next compounds were added to the first row of a 96-well plate. Media with DMSO was then added to the rest of the rows. The compounds were then serially diluted row wise and the last row was designated as the negative control row with just DMSO added. These conditions were added to the spheroids to induce apoptosis.
The results demonstrated that a full plate could be treated in less than 7 minutes. One key area that was addressed was that the tip position and speed of aspiration had to be strictly controlled to ensure no drops were lost and no spheroids were aspirated. Also the addition of DMSO caused droplet instability due to reduced surface tension, so some media had to be removed before the compound was added.
Prior to analysis on day 4 the spheroids were checked for consistency across wells, which is critical in compound screening. After being analyzed, it was determined that the spheroids showed consistent size and circularity.
Day 4
On Day 4, samples were prepared for analysis. Analysis methods included high-content imaging using ImageXpress Micro from Molecular Devices and flow cytometry conducted using the Gallios by Beckman Coulter.
High-content Imaging
Sample preparation for high-content imaging required that 6 μl of media be removed and 6 μl of diluted staining reagent be added. A full plate was processed in under 8 minutes and then was placed in the incubator for two hours.
Imaging 3D cultures presents several challenges compared with 2D adherent culture. In 2D adherent cultures high magnification images can be obtained because the culture is flat, has a single plane of focus and is stationary. With hanging drops, there is no single plane of focus and the spheroids can move around within the drop, thus eliminating taking multiple images and then assembling. As a result, lower magnification was required to capture the full sphere.
Goals for automation of the imaging portion were to minimize dead volume as stains can be expensive, no lost wells due to the pipetting or movement, no lost spheres during the media change and the ability to image the sphere at 10X.
To meet these goals and address 3D imaging challenges, the ImageXpress Micro, was used to capture fluorescent images of the spheroids after compound treatment. Images were captured at 10X magnification and pauses between filter changes enabled images to be taken while the spheroid was stationary. This allowed Z-stacks to be taken and therefore guaranteed the optimal plane of focus.
As expressed in the webinar, there were lessons learned in the automated imaging process these included:
- There was quite a bit of evaporation that caused the spheres to “ride up” the droplet, adding water in the plate reservoirs helped to mitigate this.
- It took Z stacks to improve image focus
- Diffusion into sphere varied with reagent
- Some compounds were fluorescent and stain removal would be difficult
Flow Cytometry
Spheroids were transferred to a 96-well round bottom plate containing PBS. Then PBS was removed and Accumax was added to the wells and using the 96-channel EST head to mix and dissociate the spheroids. Next staining reagents were added to cytometry tubes and cells were relocated to flow cytometry tubes. 96 wells were processed in 48 minutes, stained and then incubated for 1 hour.
There were also challenges associated in obtaining cellular data from the 3D cultures. In suspension cultures, cellular data including size and fluorescence is simpler as you can collect thousands of data points per sample and essentially look at one cell at a time. With the 3D culture spheroids, they needed to be dissociated, which required a pipetting technique to transfer the spheroids and programming for spheroid dissociation.
Goals for automation of the flow cytometry portion were that all spheres would be transferred, complete sphere dissociation would occur and successful transfer of the cells to flow cytometry tubes. These goals were achieved.
There were also lessons learned in flow cytometry that included:
- Dissociation was more challenging with older spheres (i.e. day 7), there was more cell death (i.e. higher concentrations)
- Dose responses could differ from imaging
Complimentary Analysis
By using two methods of analyzing the culture, a more complete picture of the impact of the compounds was reached.
Study Takeaways
The study presented in this webinar, successfully demonstrated the following:
- Automated cell plating and treatment provided excellent consistency in cancer spheroid formation while minimizing the loss of hanging drops.
- Automated sample prep enabled sphere-level analysis via high content imaging and cell-level analysis via flow cytometry
- The combination of these two analyses provided a more complete understanding of the response of cancer spheroids to apoptosis inducers
Looking forward

By successfully demonstrating that 3D culture can be plated, treated and analyzed through automation it confirms that this process can be scaled up for a full compound screen that would involve multiple plates. Plate and tip storage and an integrated analyzer would need to be added. In addition, it would be essential to have software that could enable a multi-plate screen, this would include optimizing the labware movement, multi-day scheduling, data management, remote monitoring and a process where results could drive pipetting. A benefit of the Biomek FXP is that it can be customized for different integrations both on deck and in adding instruments including analyzers, incubators, barcode readers, and washers.
Conclusions
It is clear that 3D culture provides a more accurate method for conducting research into cell-to-cell interactions and drug compound screening. However, the path to implementing this system can be daunting and unclear. Automated systems including automated plating solutions and automated workstations like the Biomek FXP solves many hurdles that previously made implementation of 3D culture too labor intensive or not high throughput enough. While implementing an automated system may still feel a bit overwhelming, partnering with companies like Beckman Coulter or others in this area can help you create a system custom designed to achieve your goals.
For more information about Beckman Coulter’s automation of 3D processes please see:
Webinar – Automated 3D Cell Culture and Screening by Imaging and Flow Cytometry
Application Note – Automated 3D Cell Culture and Screening by Imaging and Flow Cytometry
Poster – Automated 3D Cell Culture and Screening by Imaging and Flow Cytometry