
Gene Optimization and Synthesis as Generic Tools for Streamlined Workflows and Expression Enhancement
A Guest Blog by Dr. Arnd Dankesreiter, Sr. Product Manager, Life Technologies and
Dr. Bernd Merkl, Director of Product Management & Synthetic Biology, Life Technologies
Successful gene cloning and clone selection are key elements of creating and selecting a CHO cell line for biomanufacturing. Traditional cloning methods involve many steps from sequence to clone selection and offer little control over the final gene. Since the process is imprecise, often the resulting gene is of poor quality, has mutations, or produces no gene at all.
Today’s de novo gene synthesis technologies offer easy, fast, and affordable access to nearly any desired gene sequence. A viable alternative to cumbersome cloning and mutagenesis steps carried out in individual labs, gene synthesis offers researchers maximum design flexibility and makes it possible to receive a 100% correct clone in around 5 business days. In addition, this technology makes every sequence available to scientists and offers the ability to utilize the redundancy of the genetic code to optimize protein coding genes and open reading frames of choice for maximum protein expression. Gene optimization and synthesis services offer an attractive solution for cloning needs. For example, Life Technologies GeneArt® has a capacity of 5.5 Mbp, which enables researchers to simplify their workflow, free internal lab resources for value-added work, and to gain speed in research and development by increasing the success rate based of gene expression projects.
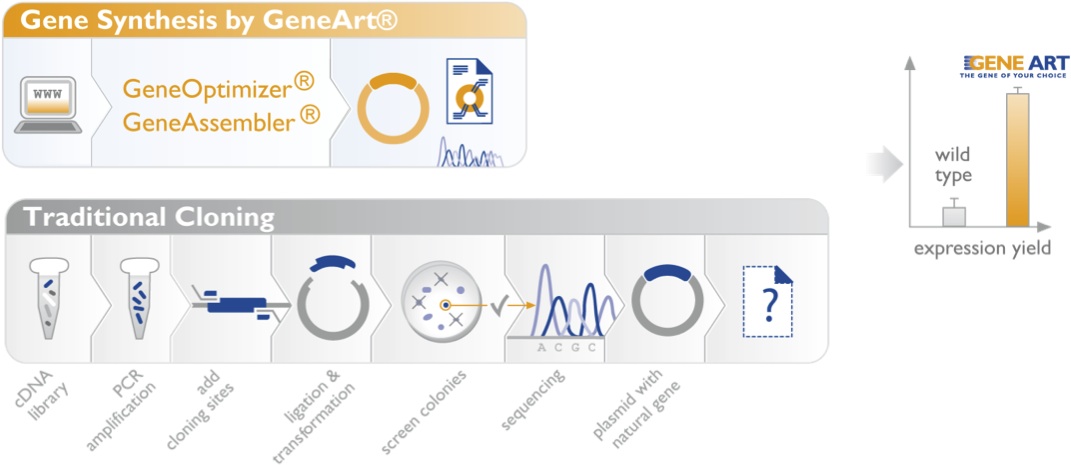
Going Beyond Codon Optimization for Better Expression
For synthesized genes, expression success depends largely on the quality of the optimization algorithm. The algorithms must take into account the major contributors to high-level gene expression, including genetic stability, transcriptional efficacy, and mRNA stability. This helps ensure that the resulting synthesized gene is fully optimized for expression in the host organism. Much more than codon optimization gene optimization is a true multi-parameter challenge.
Success is Demonstrated in the Expression of Optimized Genes
When evaluating the success of a gene synthesis program, expression must be the key parameter for success. Quality of the algorithm must be tested and verified by confirming that the optimized gene candidates were able to express and that expression levels were increased when compared with their natural counterparts. In addition, gene optimization is not only a tool to make expression success rates more predictable and improve expression yields in general, but also optimized genes represent a powerful tool in functional genomics as the sequence-modified counterpart can be used for the rescue of an siRNA-mediated knockdown as demonstrated in Fath et al.
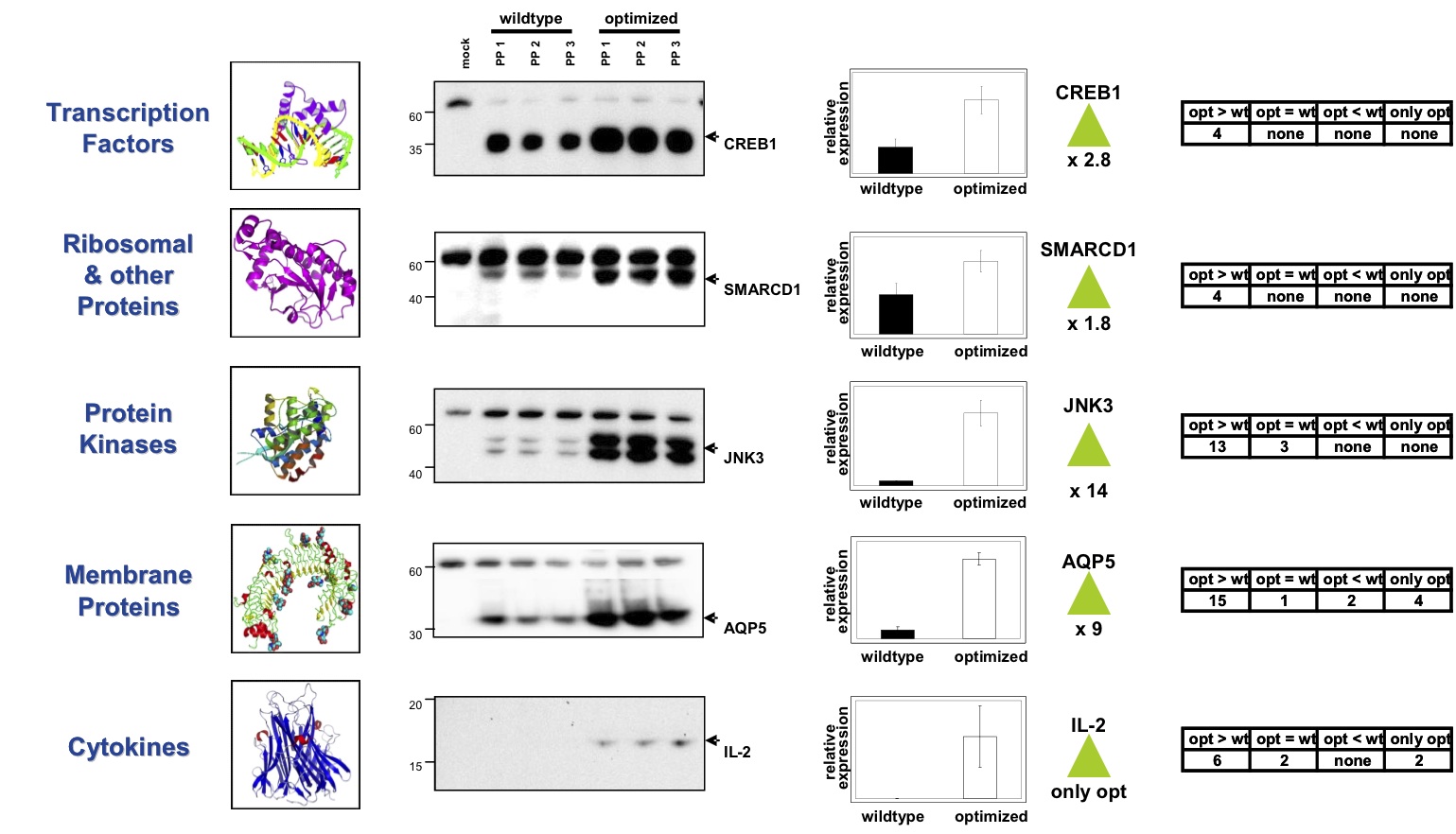
Life Technologies Gene Optimizer Technology
The Life Technologies GeneOptimizer® software suit has been developed over the last 15 years by extensive literature research and parallel lab verification. To systematically test the quality of their algorithm, 50 GeneOptimizer® RNA and codon optimized candidate genes representing five classes of human proteins—transcription factors, ribosomal and polymerase subunits, protein kinases, membrane proteins and immunmodulators—have been compared with natural genes in autologous expression in HEK293T cells.
The results show that GeneOptimizer® algorithm increases the overall expression success rate: 100% of the optimized genes did express and 12% of the natural genes did not express in HEK293T cells. Furthermore, 86% of the optimized genes showed higher expression than their natural counterparts, while 10% expressed at the same level in the autologous system. Only 4% showed lower expression than their natural counterparts. These were mainly genes coding for membrane proteins, and lower expression may be expected in such proteins as cell toxicity caused by overexpression is not uncommon. The full results have been published: Fath et al. (2011), PLoS ONE.
The protein expression increases offered by GeneOptimizer® technology have not only been proven in comparative studies with natural genes but also when compared with gene optimization algorithms from other suppliers of synthesized genes. As seen in Figure 4,, the GeneOptimizer® optimized gene showed a 3-fold increase compared to the natural gene, while the genes optimized by algorithms from other companies resulted in protein expression that was lower than the natural gene.
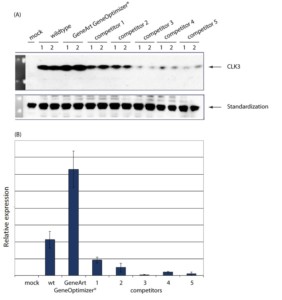
Comparative expression analysis of wildtype versus gene-optimized constructs:
(A) Western Blot analysis using a-His antibodies. Expression levels of 2 independent transfections per wildtype and optimized construct are displayed. Articlesization is based on endogenous a-His protein.
(B) Summary: Relative expression levels of wildtype versus optimized constructs of three independent transfections are displayed.