
Modernize Your Cell Development Assays with High-Throughput Image Cytometry
Cell line development encompasses an optimized process that balances cell line productivity and cell product quality over a consistent and reproducible timeline. This article summarizes areas of cell line development where image cytometry has provided workflow advantages and made the entire process more efficient.
There are several solutions available for research groups within cell line development. For most manufacturers, CROs, and larger scale operations, there can be a daily need to process stacks of plates utilizing a high-throughput solution. As bioprocessing moves towards high-throughput and automation solutions – it seems prudent to replicate already established methodology from leaders in cell line generation.
CLD Workflow Automation
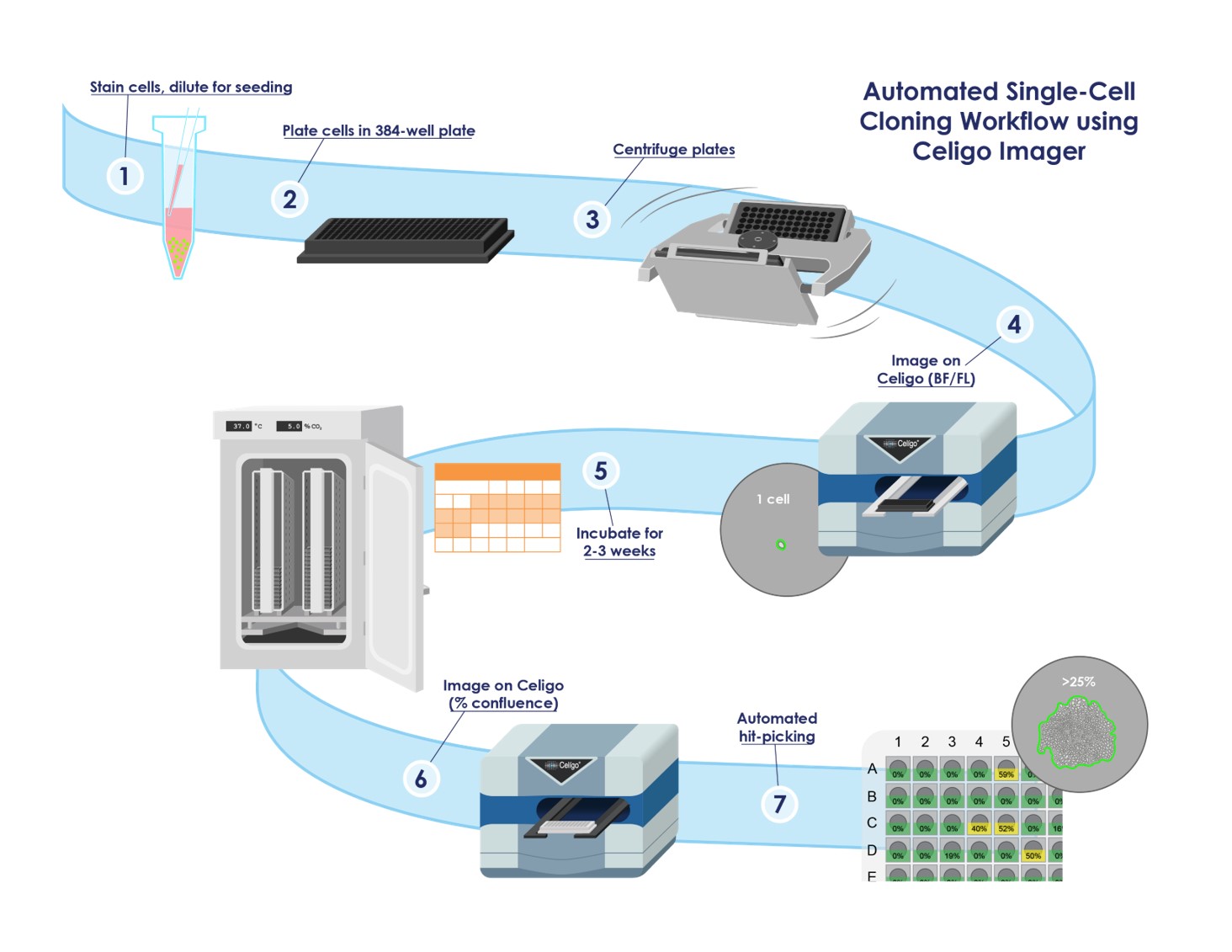
Though single-cell cloning is a hallmark of the cell line development community, the entire process of engineering a cell line is composed of multiple stages (Figure 1). Genentech, Inc. is often attributed to originating and continuing to pioneer methodology that shapes cell line development. Recent publications disclose Genentech’s automated single-cell cloning workflow with implementation of imaging plates at the time of seeding to immediately identify wells likely to containing single clones.1, 2, 3 This process was optimized by fluorescent live-cell staining with CMFDA, which increased accuracy of identifying single cells within wells. Plates were imaged in brightfield and fluorescence, and growth was tracked until confluent wells could be picked. Monoclonality was visually confirmed prior to banking of any cell lines. Genentech utilized Nexcelom Bioscience’s Celigo Image Cytometer within their automation workflow.1, 2, 3
Fluorescent Labeling
Though debated within the cell line development community, authenticity of single-cell clones can be enhanced with the use of a fluorescent marker. The WHO Expert Committee on Biological Standardization (ECBS) have stated that any changes to the cell culture process should not affect product quality4,. Therefore, fluorescent labeling should be limited to reagents that will not interfere with the integrity of the cell line. The only tracer dyes that should be used are transient ones, unless a cell line has been engineered to fluoresce for non-clinical research applications. Genentech has thoroughly vetted the safety and integrity of utilizing CMFDA for verifying clonality1. CMFDA is a green cell tracer dye that is transiently retained inside live cells. CMFDA is a cell-permeable compound that is rapidly hydrolyzed inside live cells converting it to a fluorescent marker. The dye becomes difficult to detect after approximately 3-5 generations. Studies strongly suggest that the use of CMFDA during single-cell cloning does not interfere with safety or overall product quality, but rather reinforces the assurance of monoclonality. Ready-to-use CMFDA solution (avoid errors during resuspension and tedious calculations) for direct use on an image cytometer or for another fluorescent application can be obtained from Nexcelom Bioscience (ViaStain™ CMFDA).
Single-Cell to Single-Colony Verification
Without image cytometry, limiting dilutions are performed until the point at which, statistically, one cell should be present per well – though still not guaranteed with possible product contamination. In single-cell cloning of hybridomas, for example, it is vital to ensure that a single product of interest is generated; inclusion of more than one hybridoma cell per well could lead to the production of non-monoclonal antibodies. Working with cell banks that have a high assurance of monoclonality can help avoid serious disruptions in manufacturing. Image cytometry allows for early evaluation of clones with quantitative information that the manufacturer can use to select the most promising clones for scaling. Tracking proliferation rates and phenotypic stability of single-cell colonies through image cytometry can provide early indications for quality of the final products.
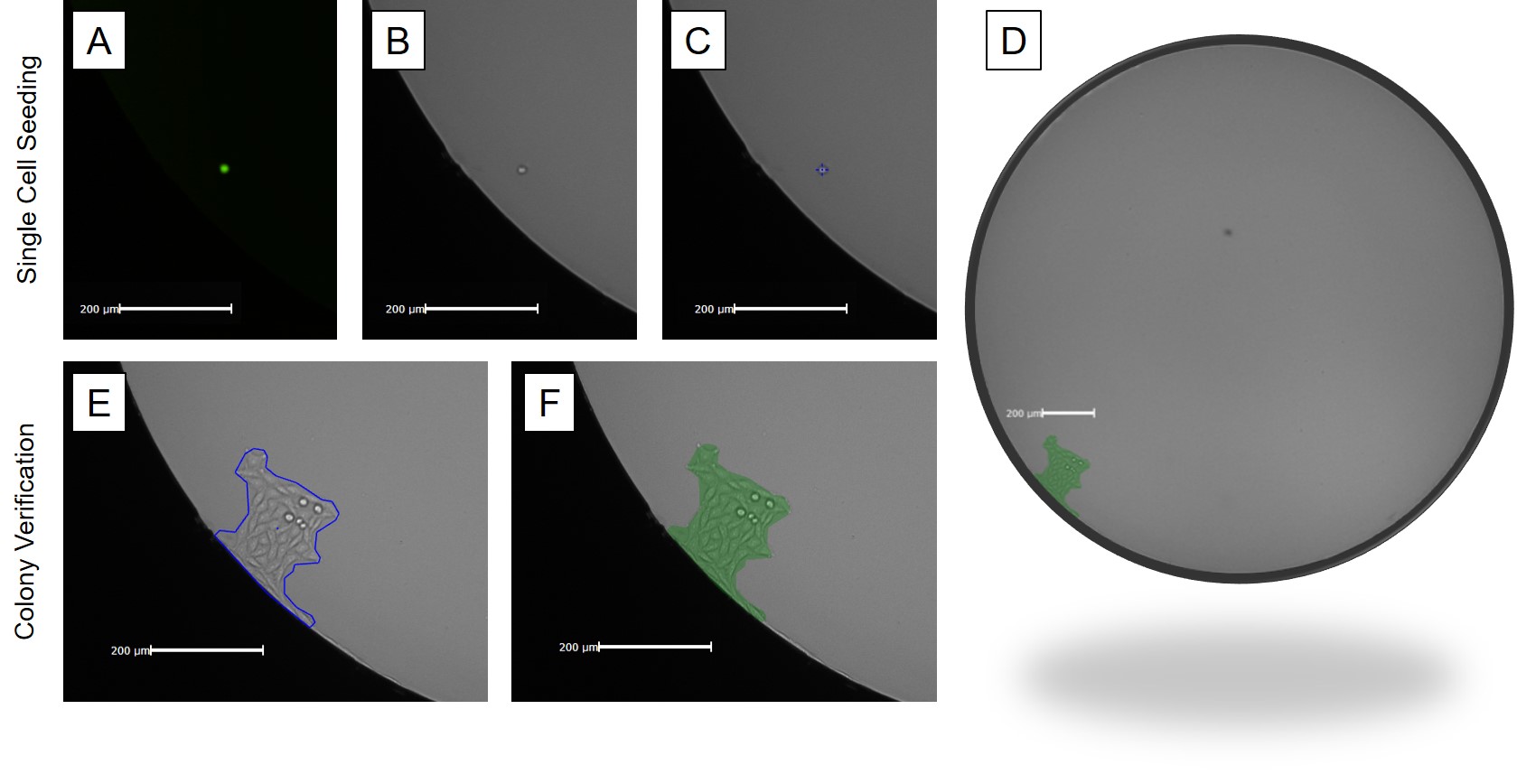
In automation, it is vital to have an image cytometer that can perform rapid whole-well imaging – especially when tracking growth of a single-cell to colony in larger well formats. In 96-well or larger area wells, poor image contrast near the edge can prevent detection of cells especially since cells often settle near the edge of a well. With the galvanometer mirrors and F-theta lens, the Celigo Image Cytometer was designed to perform rapid whole-well imaging with excellent edge contrast. With whole-well imaging, single-cells sitting near the edge of a well can be easily detected in both brightfield and fluorescent channels (Figure 2). The Celigo is frequently used as a powerful tool that provides documentation that meet FDA requirements in a single round of cloning and expansion.
Rapid and Accurate Analysis
A common goal within the cell line development community is to transition processes to high-throughput instruments that can efficiently scale up productivity without sacrificing validation. The Celigo Image Cytometer has served within this capacity as a reliable screening tool within the cell line development community. Current configurations of the Celigo allow for incredibly fast scan and analysis times that are used in automation to capture images for manual inspection over long periods of time. Genentech has published articles that demonstrate this ability by setting up projects that screen over 45,000 samples while utilizing low volume 384-well Corning 3542 plates, which are ideal for miniaturizing single-cell cloning assays.3 This method that Genentech has published allows for multi-channel scan times of less than 3 minutes per 384-well plate; it is possible to comfortably process over 5,000 samples per hour in automation. Other cell line development groups also utilize multiple Celigo instruments, some in automation, to function in cloning and other supportive projects.
Cell Printing and Imaging
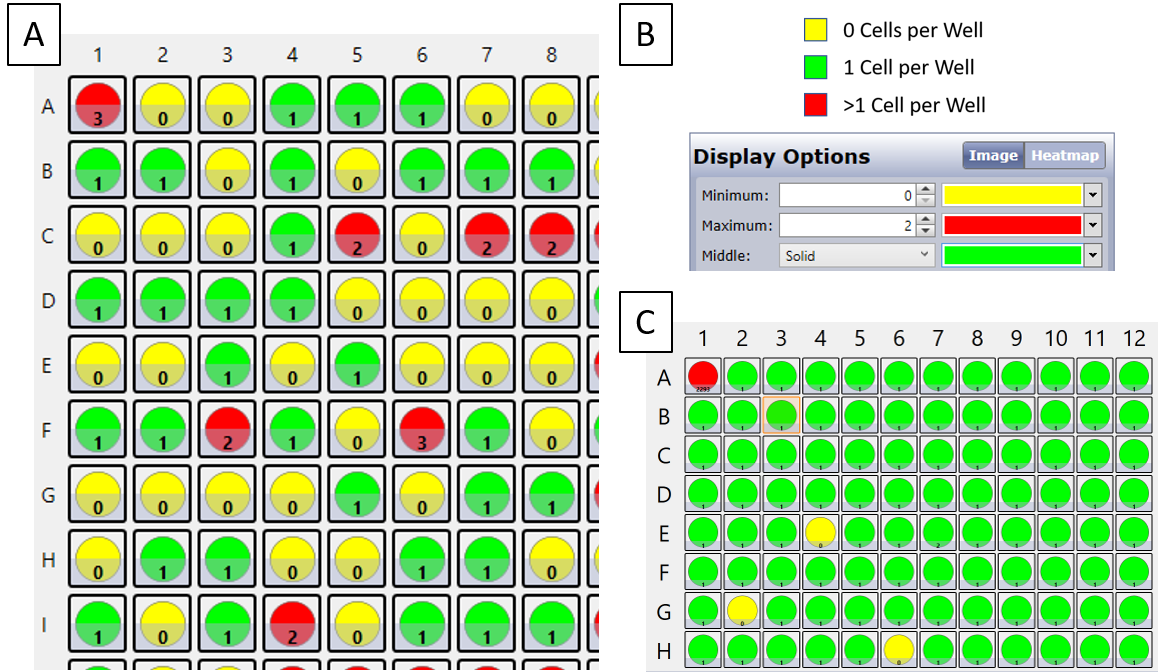
Limiting dilution can produce reliable single clones, however the entire process generates a low percentage of successful single clones per attempt. Cell printing or sorting a single cell into a well is the ideal workflow that reduces the number of plates used as well as the number of raw high-resolution images. In addition to raw images, the Celigo Image Cytometer can generate heatmaps to immediately provide the user with confirmation of seeding density (Figure 3)5. Genentech has demonstrated that combining manual image verification following image analysis from two instruments, one from single-cell printing and the other analysis tracked on the Celigo Image Cytometer, can yield an assurance of 99.9% that the clonal cell lines are derived from single cells.1, 2
Early Screening Tool
A recent US patent application by Selexis S.A. (US 2020/0109421 A1) reveals the significant role image cytometry has played in early screening of cells for production of recombinant therapeutic proteins.6 After transfection and selection, Selexis labels their cells with CMFDA followed by imaging on the Celigo platform – the same transient tracer dye and protocol used by Genentech to verify monoclonality. The patent also presents a unique use of the Celigo Image Cytometer, to detect the vitality of the cell lines being generated through a secretion application. The cell secretion application within the Celigo allows for evaluation of protein production by cells at a significantly earlier stage than conventionally practiced.7 Screening methodologies employing image cytometry can contrast antibody binding signals directly from living cells and avoid any nonspecific signals. As a high-throughput image cytometer, the Celigo has been implemented as a highly sensitive and versatile tool for hybridoma screening, capable of detecting antibody production at concentrations as low as 5 ng/mL.8 The Celigo continues to be the instrument of choice for hybridoma screening directly because of its ability to process large volumes of samples rapidly in an automated fashion; the Celigo has even been used by the FDA to study hybridomas.9 Early implementation of high-throughput screening through image cytometry can greatly facilitate antibody and therapeutic discovery.
Support for CRISPR/Cas9 Editing
Image Cytometry is frequently used in automation to track growth rates prior to batch clone screening and mass production of therapeutic products. Image cytometry has also been integral to several other aspects of cell line development especially in the generation of CRISPR/Cas9 edited cell lines. The Novo Nordisk Foundation Center for Biosustainability has published several articles about the use of the Celigo Image Cytometer in their genome technology.10, 11 Some of these applications include measuring transfection efficiency rates for mCherry-positive cells within their fluorescent enrichment protocols, cell cycle analysis with G2/M cell cycle arrest to test effects on CRISPR/Cas9-mediated genome editing in CHO cells, and to validate maintenance of cell viability. Examining cell health, including apoptosis rates and reactive oxidative stress, is also a common application for image cytometry in cell line development.12, 13, 14, 15 Overall, the Celigo can help validate and accelerate several processes, while allowing for higher throughput evaluation of targeted genes aimed at enhancing therapeutic and antibody production.
Powerful Instrument for High-Throughput Cell Counting
Many research sites have also started implementing Nexcelom Bioscience’s new high-throughput cell counter, the Cellaca MX, into various stages of cell line engineering including processing, production, and manufacturing. The Cellaca is an adaptation of existing single-chamber technologies into a format that allows for automation-level processing power at incredible speeds, 24 samples can be counted in 48 seconds with Trypan Blue or in 3 minutes with florescence. Researchers are relieving existing counting bottlenecks and are easily processing hundreds of samples within minutes instead of hours; decade-long Vi-Cell users have begun switching over to Cellaca as the next-generation solution for screening and monitoring cell health. One advantage the Cellaca provides over any traditional image cytometer is its ability to automatically load precise volumes into designated imaging chambers. Normalized volumes generate rapid yet accurate counts with significantly lower variation compared to other counting methods.
Regulatory Considerations
Regulatory processes for novel therapeutics continue to emphasize the importance of single-cell cloning and proof of monoclonality before generating a master cell bank (MCB). The FDA has identified clonality as one of the most crucial steps in guaranteeing quality and safety. The FDA recommends that two rounds of limiting dilution or FACS be used along with imaging to supplement the clonality data. The use of these additional verification steps should greatly increase the probability of monoclonality. The WHO ECBS has also stated that cloning procedures should be fully documented with appropriate statistics and detailed images.4 Assurance of clonality is a crucial part of the overall control strategy to ensure product safety, stability, quality, and efficacy. Reviewers do an assessment of the clonality of the MCB during the Investigational New Drug application stage. This review includes information on the cloning process, clone expansion, selection, and clone testing in accordance with ICH Q5D and EMA/CHMP. The FDA also requests that assurance of clonality be submitted along with the Biologics License Application. Image cytometry can be useful in supporting each of the steps within this process.
High-Throughput Solution
The Celigo Image Cytometer provides fast, time-saving, high-throughput analysis and imaging with the ability to document monoclonality and expansion with great ease. The Celigo is the “Swiss Army knife for cell line generation,” states Jean Qiu, founder and current CTO of Nexcelom Bioscience; in and out of automation, the Celigo has an extensive track record of successfully aiding in single-cell cloning, growth tracking, phenotypic stability, scale-up projects, and providing supporting documentation to meet FDA, ICH, and WHO standards. Nexcelom Bioscience is committed to supporting the Cell Line Development community by continuing to provide complete image cytometry solutions to advance research and therapeutics.
Additional resources and information can be found on these Nexcelom Bioscience webpages:
- https://www.nexcelom.com/
- https://www.nexcelom.com/applications/celigo/high-quality-imaging-and-cell-quantification-for-cell-line-development/
About the Author
 Dr Qazi received his PhD in Biomedical Engineering from CCNY/CUNY while developing and testing novel models for invasion and metastasis within the dynamic tumor microenvironment. His diverse postdoctoral research allowed him to collaborate with multiple labs within Boston (MGH, HMS), New York (MSKCC, CCNY), and San Diego (VA, UCSD). As a Field Application Scientist at Nexcelom, he focuses on sharing current research and offering better tools for advancing therapy and improving quality of life.
Dr Qazi received his PhD in Biomedical Engineering from CCNY/CUNY while developing and testing novel models for invasion and metastasis within the dynamic tumor microenvironment. His diverse postdoctoral research allowed him to collaborate with multiple labs within Boston (MGH, HMS), New York (MSKCC, CCNY), and San Diego (VA, UCSD). As a Field Application Scientist at Nexcelom, he focuses on sharing current research and offering better tools for advancing therapy and improving quality of life.
Footnotes
-
1. Shaw et al. Development and Characterization of an Automated Imaging Workflow to Generate Clonally-Derived Cell Lines for Therapeutic Proteins. Biotechnol Prog. 2018 May;34(3):584-592
-
2. Yim et al. Achieving Greater Efficiency and Higher Confidence in Single-Cell Cloning by Combining Cell Printing and Plate Imaging Technologies. Biotechnol Prog. 2018 Nov;34(6):1454-1459
-
3. Zhou et al. Beating the Odds: The Poisson Distribution of All Input Cells during Limiting Dilution Grossly Underestimates Whether a Cell Line Is Clonally-Derived or Not. Biotechnol Prog. 2018 May;34(3):559-569
-
4. World Health Organization. Recommendations for the evaluation of animal cell cultures as substrates for the manufacture of biological medicinal products and for the characterization of cell banks. WHO Expert Committee on Biological Standardization: sixty-first report; Annex 3. WHO Technical Report Series, No. 978, 2013
-
5. Zigon et al. A Rapid Single Cell Sorting Verification Method Using Plate-Based Image Cytometry. Cytometry A. 2018 Oct;93(10):1060-1065
-
6. Fourn et al, inventors; Selexis S.A. Methods for integration of transgene DNA. United States Patent Application Publication US 2020/0109421 A1. 2020 Apr 9
-
7. Harraghy et al. Epigenetic regulatory elements: Recent advances in understanding their mode of action and use for recombinant protein production in mammalian cells. Biotechnol J. 2015 Jul;10(7):967-78
-
8. Zhang et al. Novel high-throughput cell-based hybridoma screening methodology using the Celigo Image Cytometer. J Immunol Methods 2017 Aug;447:23-30
-
9. Hsu et al. Trans-acting oligodeoxythymidine phosphorothioate triester reagents for transient transfection optimized and facilitated by a high-throughput microbioreactor system. Biotechnol Appl Biochem. 2018 May;65(3):467-475
-
10. Lee et al. Accelerated Homology-Directed Targeted Integration of Transgenes in Chinese Hamster Ovary Cells Via CRISPR/Cas9 and Fluorescent Enrichment. Biotechnol Bioeng. 2016 Nov;113(11):2518-23
-
11. Grav et al. One-step generation of triple knockout CHO cell lines using CRISPR/Cas9 and fluorescent enrichment. Biotechnol J. 2015 Sep;10(9):1446-56
-
12. Lund et al. A Versatile System for USER Cloning-Based Assembly of Expression Vectors for Mammalian Cell Engineering. PLoS One. 2014 May 30;9(5):e96693
-
13. Ha et al. Baicalein Reduces Oxidative Stress in CHO Cell Cultures and Improves Recombinant Antibody Productivity. Biotechnol J. 2018 Mar;13(3):e1700425
-
14. Pristovsek et al. Using Titer and Titer Normalized to Confluence Are Complementary Strategies for Obtaining Chinese Hamster Ovary Cell Lines with High Volumetric Productivity of Etanercept. Biotechnol J. 2018 Mar;13(3):e1700216
-
15. Hansen et al. Versatile microscale screening platform for improving recombinant protein productivity in Chinese hamster ovary cells. Sci Rep. 2015 Dec 11;5:18016