
Achieve unparalleled structural elucidation and sensitivity with the ZenoTOF 7600 system from SCIEX
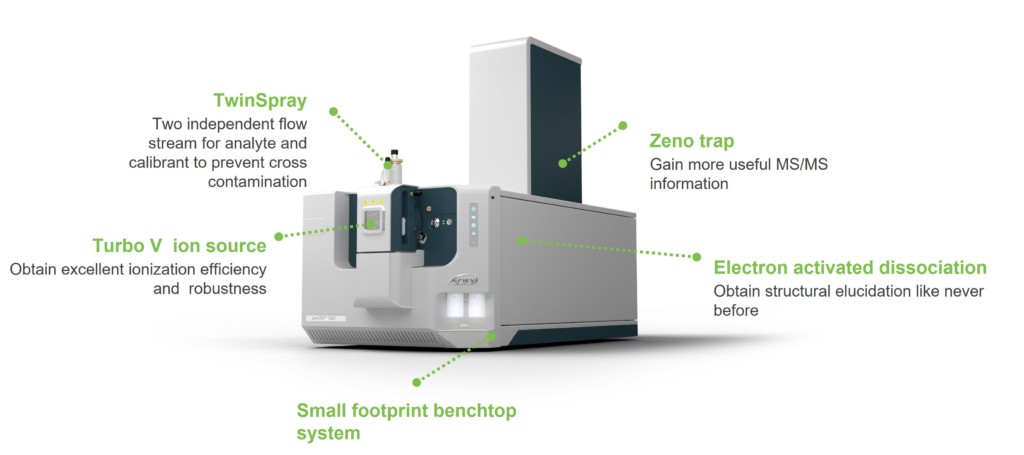
Liquid chromatography with tandem mass spectrometry (LC-MS/MS) is one of the most versatile and high-performance analytical tools for the detection, characterization and quantification of biotherapeutic molecules and vaccines. However, emerging modalities require new analytical methodologies to enable in-depth characterization and mitigate potential risks early in the development process. In addition, the increasing potency of next-generation biologics drives the need for more sensitive assays for bioanalysis studies. Uncovering new perspectives for molecules, which might have even been previously undetectable, requires overcoming the specificity and sensitivity limitations of current LC-MS/MS technologies. In response to these needs, SCIEX has launched the ZenoTOF 7600 system. This accurate mass, quadrupole time-of-flight (QTOF) system can improve compound identification in protein, oligonucleotide, metabolite and lipid analyses, and it can help scientists uncover new insights and drive decision making processes faster. In addition to offering collision-induced dissociation (CID), the instrument offers a new tunable fragmentation technology based on electrons. The tunable electron activated dissociation (EAD) technology tackles the elucidation of challenging molecules across all charge states, allowing for new depths in the characterization of molecules. Pairing either dissociation mode with the Zeno trap enables higher levels of sensitivity for unparalleled accurate mass quantification, and allows scientists to:
- Characterize large molecules such as proteins and oligonucleotides, including post-translational modifications (PTMs), with unprecedented depth
- Elucidate positional isomers in lipids and amino acid isomers in peptides
- Identify and quantify oligonucleotides, proteins and peptides with unparalleled sensitivity and speed
Gain more information on modified peptides
In the manufacturing of protein therapeutics and viral vectors for gene delivery, product and critical quality attributes such as post-translational modifications (PTMs) need to be fully characterized to ensure safe and effective biologic drugs with consistent quality. While CID is the go-to-methodology for LC-MS/MS-based identification, with its high fragmentation efficiency and excellent reproducibility, it leaves gaps with regard to some critical structural questions. Especially the exact localization of PTMs (such as N- and O-linked glycosylations, phosphorylations or sulfations) on peptides and the differentiation between amino acid isomers cannot be determined with CID. Although alternative fragmentation techniques exist, their wide adoption has been limited due to the need for tedious charge-state dependent tuning, poor fragmentation efficiencies (especially for low-charged species) and low acquisition speed, to name a few.
With the introduction of EAD, peptide analyses can be performed with new structural depths, while ensuring efficiency in generating results by using a generic platform method with one kinetic electron energy setting capable of dissociating peptides across charge states. As an example, the amino acid isomers aspartate and iso-aspartate derived from deamidation events of an adeno-associated virus (AAV) capsid protein can be differentiated with the EAD platform method (Figure 2).

EAD provides reproducible and rapid fragmentation that can be used to accurately determine PTM localization and isomer differentiation that is well-suited for routine analyses to meet the rigorous analytical requirements of monoclonal antibodies and structurally complex next generation protein therapeutics, vaccines and viral-based delivery systems.
Taking lipid nanoparticle (LNP) characterization to the next level
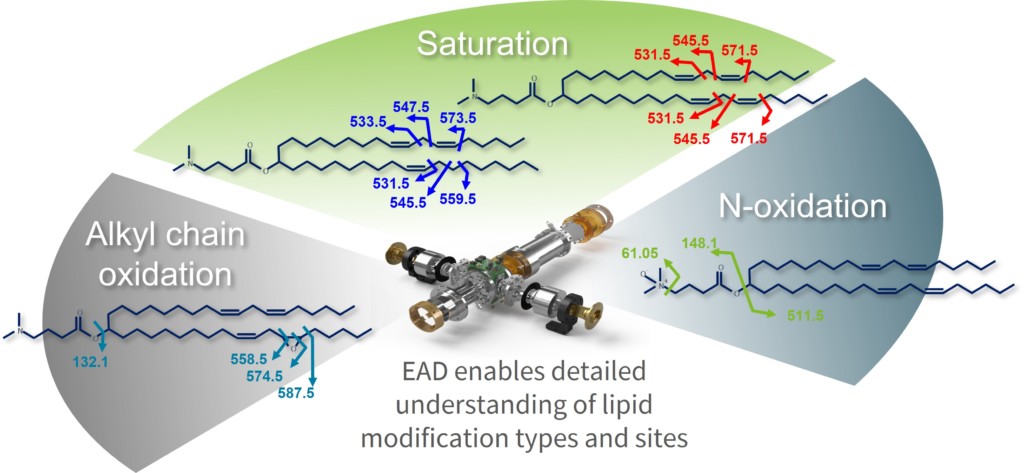
Lipid formulations are of primary importance for ensuring the stability of the encapsulated genetic material, such as oligonucleotide therapeutics or mRNA-based vaccines. Recent studies showed that modifications on lipids can negatively affect the genetic cargo. Understanding the lipid material used for LNPs with higher levels of depth can therefore mitigate risks during development of new modalities. Low-charged lipids in particular have posed an analytical challenge, since structural information gain with CID is fairly limited and alternative fragmentation techniques weren’t applicable to the frequently singly charged class of molecules. The ability to tune the kinetic energy of the EAD technology in an easy manner provides access to multiple fragmentation mechanisms that can be applied to a diverse array of modified peptides and to lipids to extract in-depth structural information, providing a unique fragmentation profile that was previously impossible to achieve (Figure 3). This is especially advantageous for the characterization of novel lipid structures to achieve complete profiling and confirmation, including fatty acid attachment points, location of double bonds, cis or trans configuration and N-oxidation.
Never miss an important analyte
Highly potent drugs, a focus on personalized medicine with lower yields and the goal of reducing animal studies all drive the need for more sensitive assays. Pairing the higher information gain from EAD with another novel technology — the Zeno trap — is the perfect match to enhance MS/MS data quality and signal-to-noise. Ions are accumulated in the Zeno trap before being rapidly pulsed into the TOF chamber, resulting in the detection of 5-20x more ions and significantly enhancing sensitivity and spectral quality (Figure 4). While pulsing the Zeno trap is particularly useful for EAD experiments, it can also be used for traditional CID to achieve the highest sensitivity.

Meeting digitization needs – To meet the needs of modern-day laboratories, the ZenoTOF 7600 system is powered by a suite of advanced software tools that can seamlessly integrate with existing data pipelines. Automated acquisition, processing, reporting and data management is achieved with SCIEX OS software. In addition, Biologics Explorer software and Molecule Profiler software further extend the analytical capabilities of the advanced hardware to streamline workflows and deliver high-quality, actionable data for a wide range of applications from proteins, peptide digests, oligonucleotides, therapeutic peptides and small molecules.
The technological advancements of the disruptive new LC-MS/MS platform combine qualitative flexibility and quantitative power, allowing scientists to identify, characterize and quantify molecules with better sensitivity, accuracy, specificity and speed than ever before. The ZenoTOF 7600 system is a breakthrough that helps scientists overcome the developability gap in bringing new biotherapeutic molecules to market faster.
For more information on the technologically advanced ZenoTOF 7600 system, please visit https://sciex.com/products/mass-spectrometers/qtof-systems/zenotof-7600-system.