
Design of Experiment Principles to Optimize Benzonase® Endonuclease in Viral Gene Therapy Applications
For over 30 years, Benzonase® endonuclease has been used for the removal of nucleic acids in virus vector and vaccine manufacturing as mandated by regulators to ensure the purity and safety of the final product. The enzyme can degrade both DNA and RNA into small 3–5 base pairs fragments (<6 kDa) with no base preference, making it a versatile tool for process development to optimize the yield in virus purification as well as to improve the efficiency of downstream chromatography and filter devices. However, Benzonase® endonuclease activity is strongly influenced by several factors (Figure 1) and balancing these is a critical step during process development to maximize its benefit in the overall workflow.
Statistical design of experiments (DoE) is a powerful strategy to help determine the relationship between factors affecting a process and the output of that process. A recent white paper, “Optimization of Benzonase® endonuclease use in virus purification,” published by Millipore Sigma provides a guide for setting up a DoE optimization experiment to achieve the optimal DNA clearance.
Setting up the DoE
In order to establish an effective DoE, it is important to define constant factors, input variables with center point starting conditions and output parameters, first (Table 1). The constant factors are key parameters that will remain constant throughout the DoE experiments. For example, the concentration of magnesium, which should be held at 2mM as it is necessary for proper function of the enzyme.
Importantly, prior to initiation of the DoE, the average DNA load of the input sample material should be determined in order to calculate the required amount of enzyme. Also note that the enzyme activity in U/μL is provided on the Certificate of Analysis for each lot and can be used to calculate the volume of Benzonase® endonuclease using the equation in Table 1, center point starting conditions. The center point conditions provide users with a point of reference from which parameters can be altered to determine the relationship between the input and output factors. Basing DoE experiments on enzyme activity, instead of protein concentration, ensures reproducible DNA digestion.
| Parameters | |
|---|---|
| Constant Factors | Average DNA load Equipment: incubation vessel and mixing process Upstream material: pH and salt concentration [Mg2+] = 2mM |
| Input Variables | Benzonase Concentration Incubation Time Incubation Temperature |
| Center Point Starting Conditions | Benzonase Concentration: Incubation Time: 2 hours Incubation Temperature: 21°C |
| Output Variables | DNA content of >100 bp fragments Product Integrity |
Table 1. Summary of the parameters important to establish DoE an effective for Benzonase® endonuclease
Ultimately, the goal is to achieve optimal DNA digestion and thus it is recommended that DNA digestion by Benzonase® endonuclease be monitored using a specific qPCR method for a 100-200 base pair fragment of host cell DNA. Because complete DNA digestion by Benzonase® endonuclease results in small 3-5 base pair fragments, this limits the utility of fluorescent DNA-intercalating dye assays that can detect DNA fragments as small as 4 base pairs. Additionally, having the appropriate control experiments to define the digestion process ensures the integrity and robustness of experimental results.
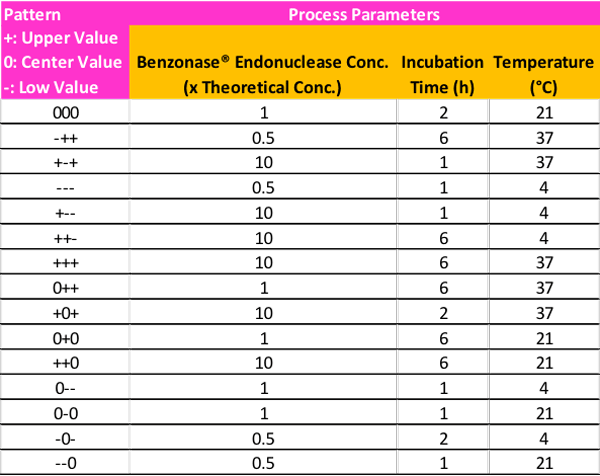
As a starting point, two values, one above, and one below the center point starting conditions, can be utilized in the optimization experiments to monitor enzyme performance as a result of changing process parameters. These can be denoted by upper- (pattern symbol +), center- (pattern symbol 0), and lower value (pattern symbol -). A potential example of the different permutations based on this guidance for the DoE matrix can be seen in Table 2.
Interpreting your DoE Results
The experimental design proposed in Table 2 is a suggestion based on the main process parameters that influence DNA/RNA digestion by Benzonase® endonuclease. There are, of course, multiple approaches for determining the set of design points to be used in the experiment, and it is also possible to increase or reduce the number of conditions depending on resources and time allotment.
Once the DoE has been executed, the data obtained should be compared with the initial experimental assumptions prior to full analysis and interpretation. The data generated may lead to additional experiments or DoE where the parameters are more narrowed and focused around the potential optimal parameter values. Additionally, optimization at the point of use, such as in the bioreactor, post-cell lysis or post-clarification should also be considered to achieve the most economic and optimal performance of Benzonase® endonuclease for the manufacturing process. This example DoE serves as a starting point to optimize operation conditions that deliver the DNA clearance required.
For more information, please see Optimization of Benzonase® endonuclease use in virus purification