More Fragments, Less Headaches: How Electron Activated Dissociation Can Improve Lipid Nanoparticle Analytics in Gene-Based Drug Development
In contrast to traditional biologics, drugs based on the delivery with lipid nanoparticles (LNPs) can be even more complex and pose new challenges from an analytical perspective. The genetic cargo and several different lipid classes and impurities must be properly characterized and quantified to ensure the safety and efficacy of the final drug product. Specifically, the lipid composition can have a significant impact on the functionality of the active ingredient, the oligonucleotide.
Solving analytical challenges of LNPs is the focus of the webinar Handle with care: Ensuring LNP lipid quality for better genetic medicines. These challenges, and their potential solutions, are presented by Adam Crowe, PhD, Manager of Analytical Development at Precision NanoSystems ULC, a company that specializes in the production of LNPs with proprietary lipid libraries, instrumentation for LNP formulation and drug development expertise, and that understands the complexities of LNP-based therapeutics and vaccines and the importance of lipid raw material analysis for LNP development and therapeutic success.
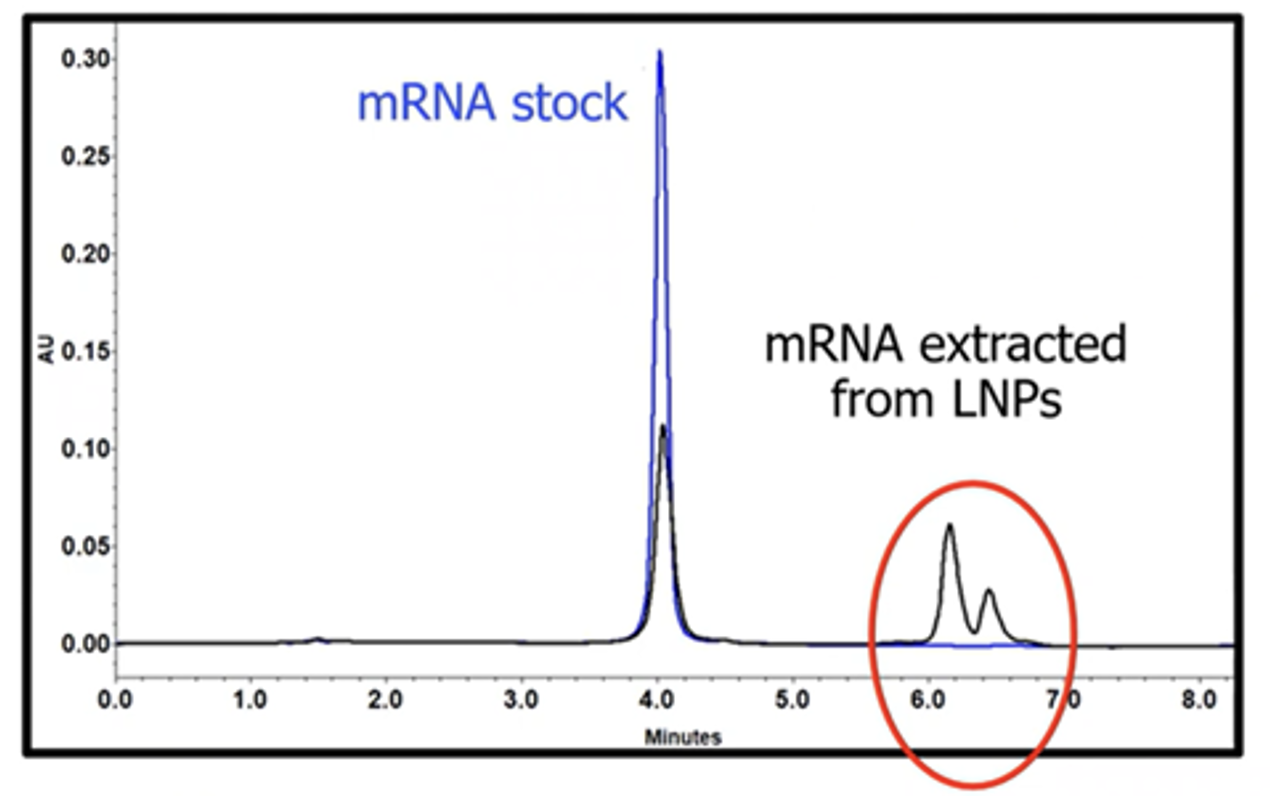
In the webinar, to demonstrate the complexity of LNP analytics, Crowe compares the chromatograms of stock messenger RNA (mRNA) and mRNA extracted from LNPs (Figure 1). While a typical chromatogram of an mRNA sample before formulation usually shows one peak (blue trace), mRNA extracted from LNPs frequently shows additional, more hydrophobic peaks (black trace). Crowe explains that, until recently, the identity of the extra elution peaks was unknown, and culprits such as aggregated mRNA or residual lipids did not make sense considering the results of other analytical experiments. Crowe’s team therefore strongly suspected that the mRNA was being modified covalently, causing it to elute separately.
Because LNP formulation is based on intermolecular interactions between the lipids and the oligonucleotides, such as mRNA, it makes sense to look at the LNP composition to determine the root cause of the additional mRNA peaks. LNPs are an analytically complex mixture consisting of four main lipid components: ionizable lipids, cholesterol, helper lipids and PEG-lipids along with their genetic cargo. The ionizable lipids exhibit a positive charge at low pH, which facilitates complexation of the negatively charged nucleic acid and encapsulation through electrostatic forces. The cholesterol, helper lipids and PEG-lipids provide stability during nanoparticle formation and facilitate internalization of the genetic payload via receptor-mediated endocytosis and endosomal escape with release into the cytoplasm of the target cells.
Ionizable Lipids Enable LNP Technologies
Ionizable lipids play a key role in nucleic acid encapsulation during nanoparticle formulation along with other aspects of nanoparticle behavior, such as cell targeting. Therefore, they are critical to the functionality of LNPs, and there is a growing number of new and proprietary ionizable lipids being investigated in the field for a wide range of applications.
Proper characterization, including structural elucidation, of these lipids is important for advancing LNP modalities in medicine. For example, in the webinar, Crowe explains that the ionizable lipid DLin-MC3-DMA (MC3), which is used in the formulation of the first FDA-approved small interfering RNA (siRNA) drug, has two major structural elements—the tertiary amine head group and the two identical alkyl tails—and that these elements determine the packing structure of the nucleic acid. During formulation at low pH, a positive charge is generated on the tertiary amine of the ionizable lipid that electrostatically interacts with the negatively charged oligonucleotides to induce nucleation events that, with the help of the other lipid species, self-assemble into nanoparticles.
While MC3 is a well-characterized ionizable lipid with only one tertiary amine, there are many proprietary lipids with significantly more complex structures that require cutting-edge technology for their structural elucidation.
The Challenge with Ionizable Lipids
Although ionizable lipids are proven to be very effective, they are susceptible to degradation, such as oxidation. Site-specific oxidation can occur at multiple parts of the ionizable lipids. Oxidation of double bonds of alkyl chains can change the three-dimensional structure and therefore impact the formulation of LNPs. Additionally, oxygen incorporation at the tertiary amine can result in the formation of N-oxides, which break down into reactive lipid species. A seminal paper by Meredith Packer et al. 1 suggests that reactive species derived from N-oxides of ionizable lipids can react with mRNA to form mRNA-lipid adducts. Significantly reduced therapeutic efficacy of the modified mRNA was observed, which fueled great concerns in the LNP field.
This paper provided clues for the answer to the chromatogram puzzle illustrated in Figure 1—the extra peaks are from the formation of mRNA-lipid adducts. Liquid chromatography mass spectrometry (LC-MS) methods can be used to look for the reactive species causing these impurities. It showed a mass shift of approximately +16 atomic mass units (amu) compared to the main species, indicating oxygen incorporation. However, there are other types of oxygen incorporation that do not pose the same risk to mRNA efficacy as N-oxides, and standard methods—such as collision-induced dissociation (CID), which generates very few diagnostic fragments for lipids—do not provide the MS/MS information required to differentiate the species.
Using Electron Activated Dissociation (EAD) to Characterize Lipids and Impurities
In the webinar, Crowe describes using a new fragmentation approach—electron activated dissociation (EAD)—to dig deeper into the structure of the impurities. Available on the ZenoTOF 7600 system from SCIEX, EAD is an alternative to CID that generates more fragments and provides more detailed structural information, even for singly charged lipid species, including positions of modifications and of double bonds. As a result, complex lipid species and their impurities can be elucidated with significantly more depth using EAD.
To evaluate EAD for the characterization of ionizable lipids, Crowe’s team investigated the main compound MC3 and low abundance impurities detected in the preparation. Figure 2 shows the MS/MS spectra acquired using CID and EAD, where a wealth of fragment ions can be observed with EAD while CID provides only limited information.
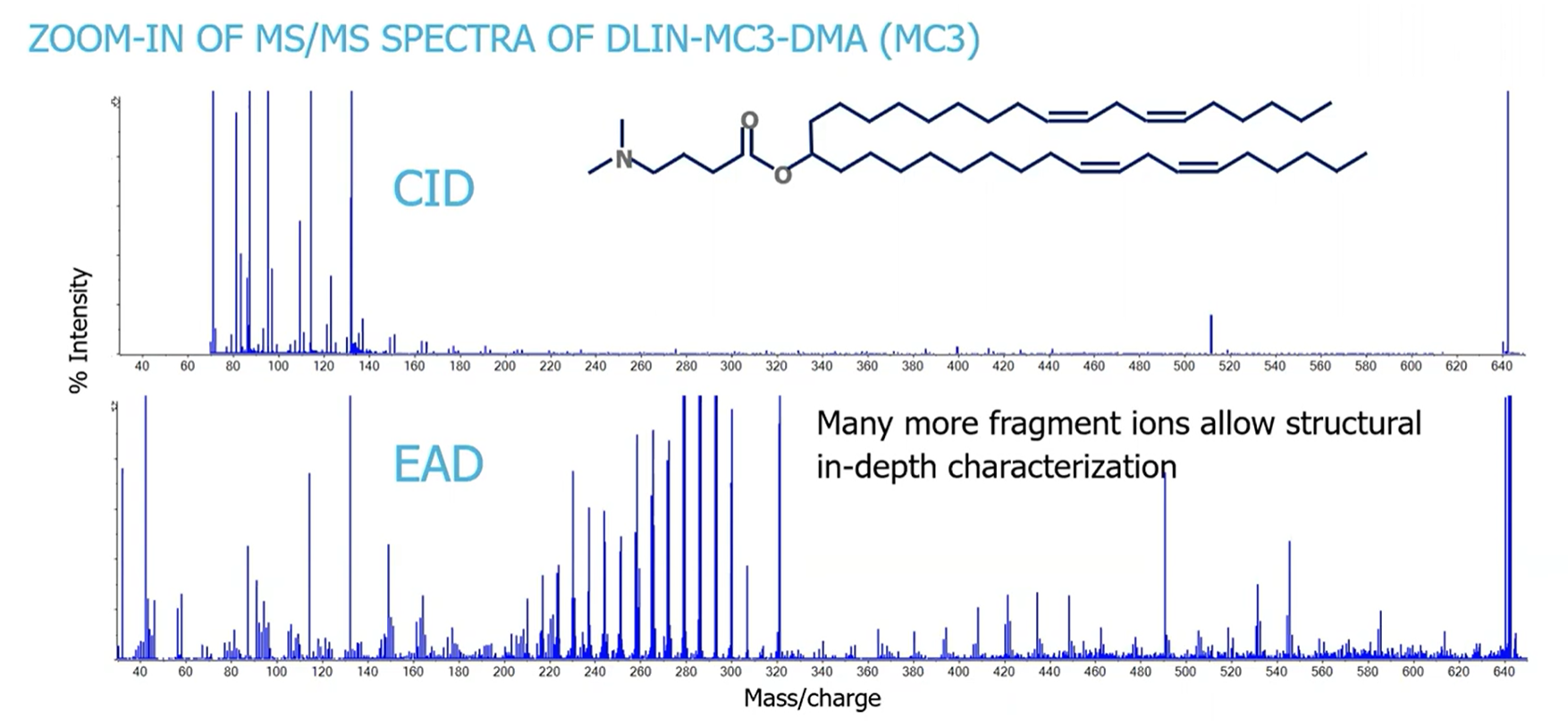
Crowe explains that every carbon-carbon bond is cleaved by EAD along the molecule, resulting in a ~14 amu shift constituting a CH2 group. Since carbon-carbon double bonds are much more difficult to fragment, minor signals with shifts of ~13 amu and more intense signals with a mass shift of ~26 amu can be observed. This allows for unambiguous localization of unsaturated carbons (double bonds). In addition, diagnostic fragments that identify the head group containing the tertiary amine could be found.
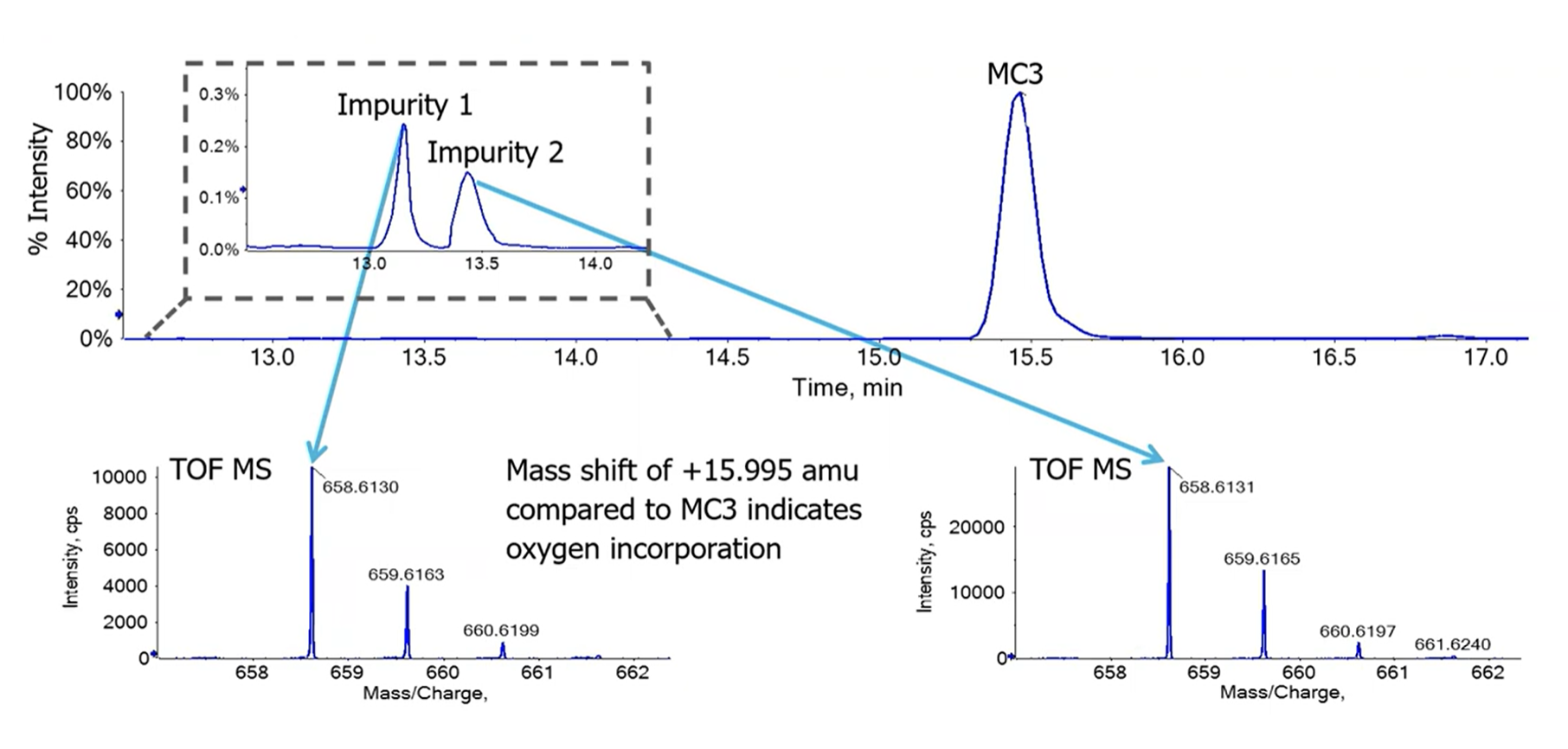
Apart from the main peak, dozens of impurities were observed in the MC3 preparation, and the highest abundant species were investigated further by Crowe’s team. Impurities 1 and 2 showed a mass increase of approximately +16 amu relative to the main MC3 species (Figure 3), indicating oxygen incorporation and the formation of an oxide species. But which species?
While the two impurities were clearly separated chromatographically, their MS spectra did not reveal any differences. When Crowe’s team applied EAD, some unique fragmentation patterns were observed between Impurity 1 and 2 (Figure 4). Impurity 1 showed the exact same fragment at ~m/z 132 that was seen in MC3, indicating the head group, but the alkyl chain fragmentation revealed unique fragment ions at ~m/z 558.5 and ~574.5, pinpointing oxygen incorporation at carbon 6 (C6). The fragmentation pattern suggested an epoxide formation at C6. From a formulation perspective, this impurity is not as concerning as N-oxides, which pose the risk of oligonucleotide adduct formation.
In contrast, for Impurity 2, the fragment of the head group at m/z 148 indicates oxygen incorporation at this site. The most diagnostic signal confirming this theory was found at m/z 61, indicating a C-N cleavage of the N-oxide species. This signal, combined with the fragments of the alkyl chains of Impurity 2 being the same as MC3, provides conclusive evidence that oxygen incorporation could only occur at the tertiary amine, resulting in the formation of the N-oxide species that is of concern.
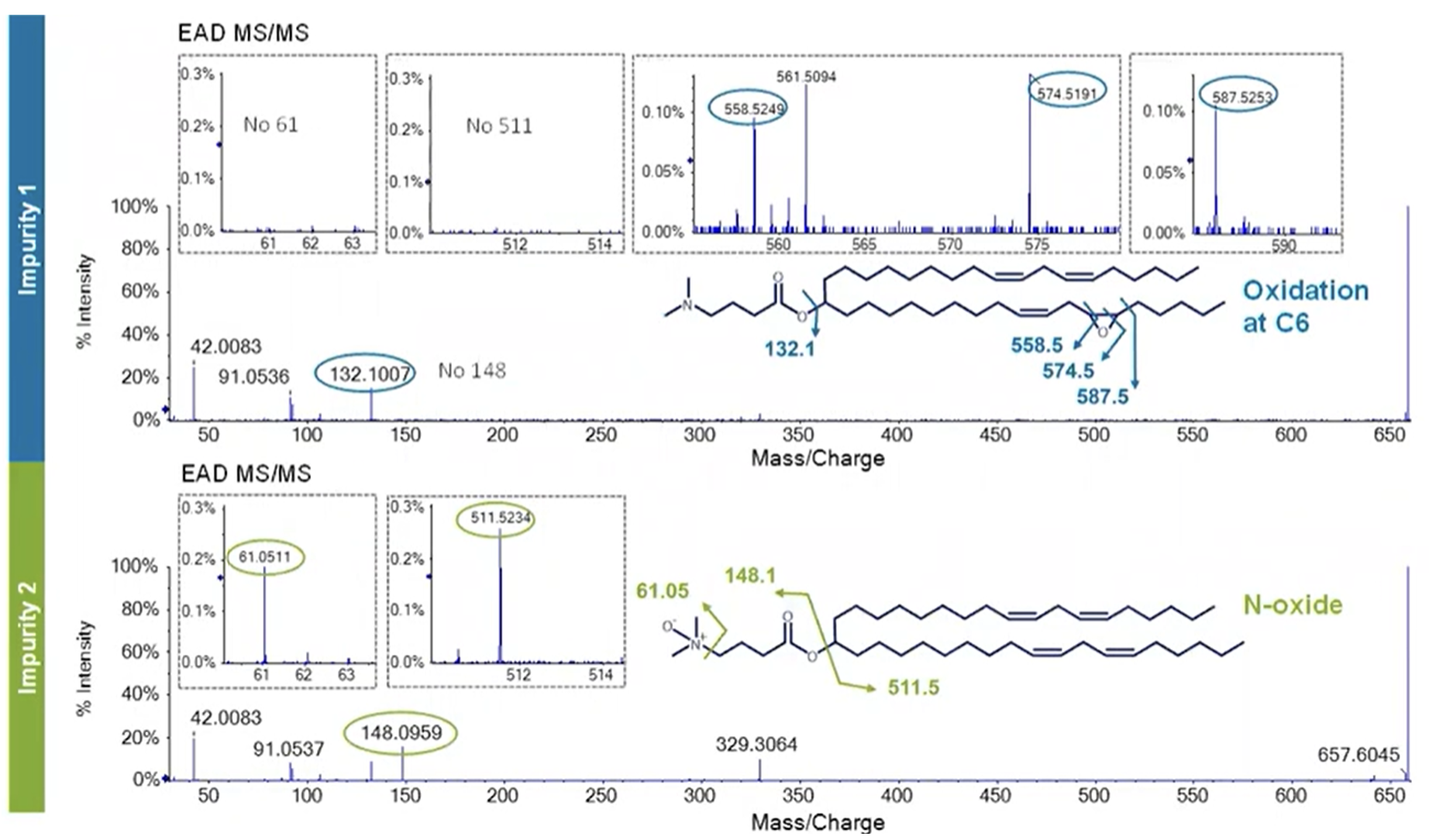
Despite the low abundance of Impurity 2—less than 0.5% relative to the main species (Figure 3)—the risk of its degradation product inactivating the encapsulated genetic material can be high. Estimates suggest that abundances of 10-5 (10 ppm) are enough to inhibit the function of mRNA, which underscores the need for detection and identification of impurities at very low abundances.
The advantage of executing EAD on the ZenoTOF 7600 system is the large dynamic range that allows signals from low abundance species to be detected leveraging a technique called the Zeno trap. TOF instruments typically transfer only 5%–25% of ions into the TOF accelerator, referred to as duty cycle. In comparison, the ZenoTOF 7600 system uses the Zeno trap, which is located between the fragmentation cell and the TOF accelerator. Ions are captured in the Zeno trap and released into the TOF accelerator with significantly enhanced duty cycle of up to 90%. Since the Zeno trap operates “on demand” automatically, a much higher dynamic range and better signal-to-noise ratio for the low abundance fragments can be achieved. This enables the structural elucidation of low abundance species based on the highest quality MS/MS data.
By using EAD to further characterize the lot of MC3, Crowe’s team was able to identify several more impurities ranging in abundance from trace amounts to 1% (Table 1).
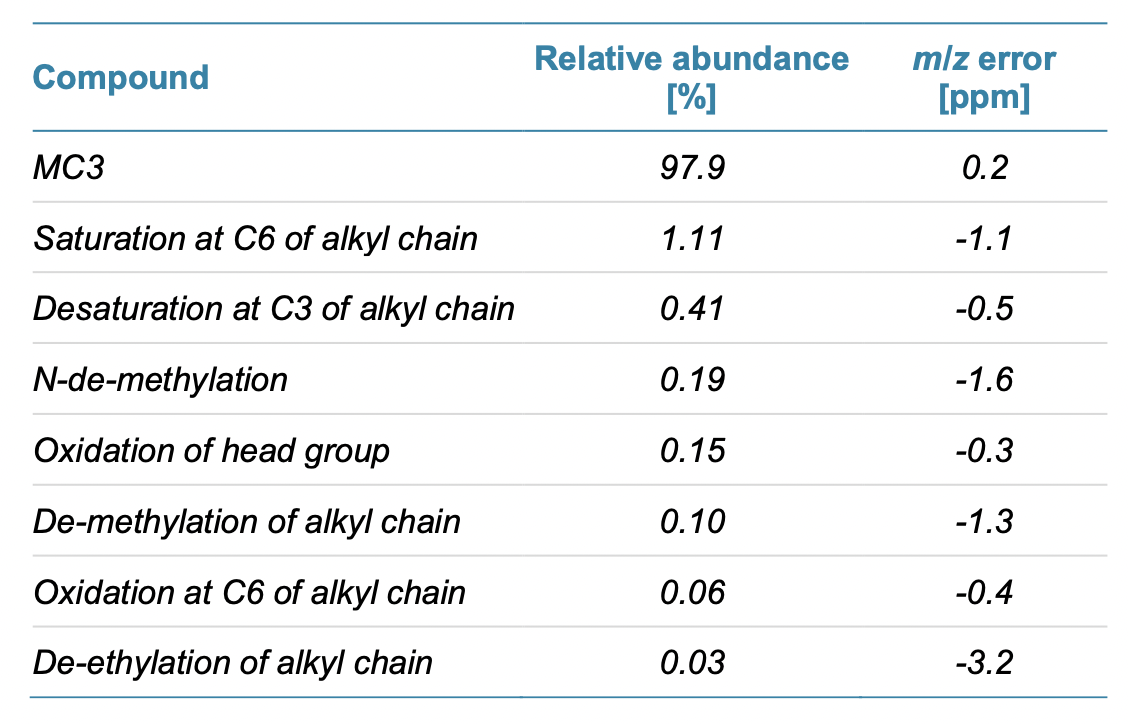
Enable Better Outcomes with Deep Characterization of Lipids
To close the webinar, Crowe emphasizes how important lipid characterization is for LNP formulation, since impurities can greatly affect mRNA activity and overall therapeutic outcomes, even at low abundance. The sensitivity and dynamic range of the ZenoTOF 7600 system can be implemented to execute this deep characterization, which is especially useful for novel ionizable lipids. The level of structural elucidation and quality of data can provide unprecedented insights into function and interactions.
According to Crowe, this is why Precision NanoSystems ULC is leveraging EAD technology on the ZenoTOF 7600 to understand lipid raw material quality and help de-risk the manufacturing program. In addition, this approach can be extended beyond ionizable lipid characterization—for example, it was shown to be effective for lipidomic applications, which greatly elevates the ability to understand lipids as a whole.
For more information, please see Handle with care: Ensuring LNP lipid quality for better genetic medicines
Footnotes
-
1. Packer, M., Gyawali, D., Yerabolu, R. et al. A novel mechanism for the loss of mRNA activity in lipid nanoparticle delivery systems. Nat Commun 12, 6777 (2021). https://doi.org/10.1038/s41467-021-26926-0